Introduction
Advances in biotechnology have enabled the development of innovative biotherapeutics, including bacteriophage-based therapies, a promising approach to combat antimicrobial resistance (AMR). However, during production, host cells express their own proteins, known as Host Cell Proteins (HCPs), which may co-purify with the final product. Even in trace amounts, HCPs can trigger immune responses, affect drug stability, and compromise efficacy. To ensure product safety, regulatory agencies such as the EMA and FDA require strict monitoring of HCP levels.
Currently, ELISA is considered the gold standard for HCP detection. However, this method has limitations: anti-HCP antibodies may not recognize all HCPs, leading to incomplete data, and ELISA provides only an overall HCP concentration without distinguishing individual proteins. In contrast, mass spectrometry (MS) offers higher specificity, enabling the identification and quantification of multiple HCPs in complex samples (1). This makes MS the preferred method in proteomics for comprehensive HCP characterization.
In this application note, ANAQUANT presents a robust method of HCP quantification using HCPprofiler, a solution designed for large dynamic range samples such as drug substances (DS) such as therapeutic phages. Given the complexity of those samples, a fractionation step is required prior to analysis to enhance detection sensitivity.
Materials and Methods
The overall strategy for therapeutic phage HCP analysis is outlined in Figure 1. Briefly, phage drug substances (DS) were denatured at high temperature under chemical conditions. The total protein content was then subjected to trypsin digestion, and the resulting peptides were fractionated under high-pH conditions using reversed-phase cartridges.
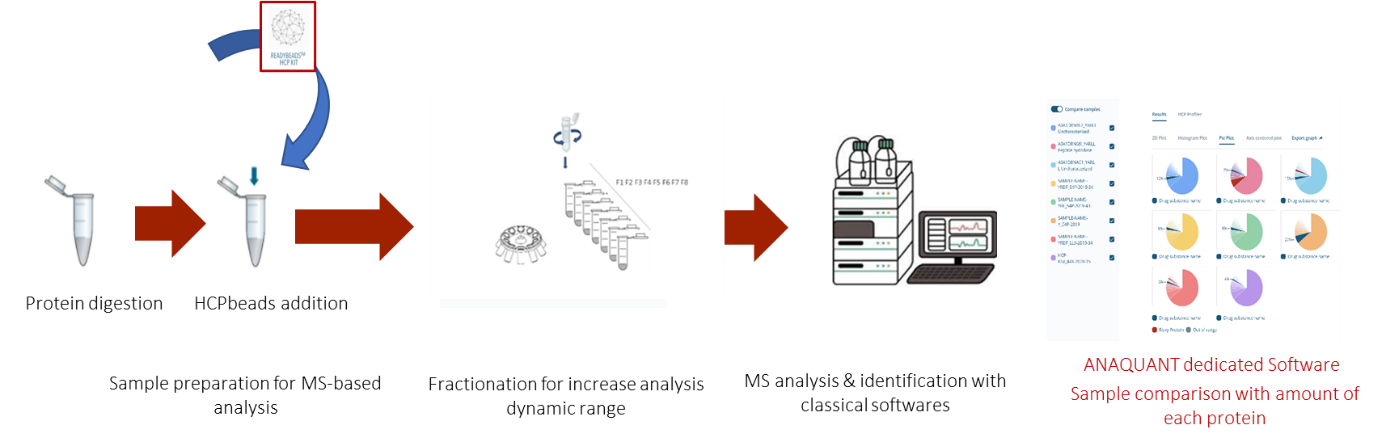
Figure 1 : Sample preparation overall workflow for HCPs identification and individual quantification in DS samples
Results
The HCPprofiler method enables both identification and individual quantification of proteins in a single experiment. HCPprofiler is also an internal quality control (QC) for analysis, assessing key parameters such as linearity (correlation coefficient R²), curve characteristics (slope and intercept), and mass spectrometer sensitivity (limit of quantification, LOQ).
Calibration curves generated across four experiments using the HCPprofiler internal QC showed strong linearity with slope around 1 and high reproducibility validating performed analysis (figure 2).

Figure 2: HCPprofiler calibration curves obtained in the four independent experiments
As expected for a therapeutic drug substance, only a small number of HCPs were detected and quantified. To further illustrate the overall method reproducibility, Table 1 presents the global quantification results for total phage proteins and HCPs. The coefficient of variation (CV%) for total HCP concentration across the four experiments was 21%, which is considered highly acceptable given the low abundance of these proteins.
HCP concentrations ranged from 0.02 to 0.54 µg/mL, with many values falling below the LOQ. These data confirm that quantification remains highly stable, even for low-abundance proteins.
| Day 1 R1 | Day 1 R2 | Day 2 R1 | Day 2 R2 | CV % | |
| Total protein concentration µg/ml | 2950* | 3370* | 3082* | 2520* | 11.8* |
| Protein numbers | 34 | 34 | 36 | 35 | |
| HCP numbers | 6 (3 out of range) | 6 (4 out of range) | 8 (7 out of range) | 5 (1 out of range) | |
| HCP protein concentration µg/ml | 1.36* | 1.03* | 1.72* | 1.58* | 21.1* |
Table 1: Global protein and HCP identification and quantification results – coefficient of variation across four replicates (*Out-of-range values included in calculation)
The DS purity evaluated using HCPprofiler in this therapeutic phage sample is 96.75% (Figure 3).
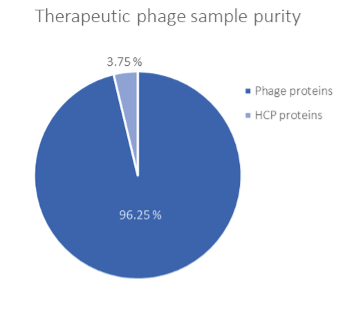
Figure 3: Therapeutic phage DS purity
This overall purity can be compared across batches over time by using the HCPprofiler solution to standardize the data.
Conclusion
Despite the structural complexity of phage capsids, the HCPprofiler method successfully enabled HCP identification and quantification in therapeutic phage samples.
To assess the robustness of the analytical workflow, we performed four independent analyses (n=4), identifying and quantifying 30 to 40 total proteins and 5 to 8 HCPs across all runs. The results demonstrated high repeatability, with consistent HCP quantification despite expected instrument fluctuations between experimental days.
These findings reinforce the robustness of our workflow, demonstrating that this approach provides accurate, repeatable, and normalized HCP quantification in complex therapeutic phage samples.
 Anaquant HCP analysis I Protein characterisation I Protein analysis
Anaquant HCP analysis I Protein characterisation I Protein analysis